Translational research
Due to the nature of our research, we are involved in various collaborations with pharmaceutical companies:
Dr. Reto Walser | Astex Pharma, Cambridge, UK |
Dr. Wolfgang Jahnke | Novartis Pharma AG, Basel, CH |
Dr. Alex Milbradt | Astrazeneca, Cambridge, UK |
Dr. Ben Davis | Vernalis, Cambridge, UK |
Dr. Alvar Gossert | Novartis, Basel, CH |
Dr. Stefan Bartoschek | Sanofi-Aventis, Frankfurt, DE |





Current Main Research Topics
Our laboratory is focusing on Drug Discovery by advanced NMR methods, including integrated methods for fast protein-ligand complex structure determination, NMR based drug design, protein allostery, and thermodynamics of protein-protein and protein-ligand interactions. We apply these techniques to therapeutically relevant receptors in-house or in collaborations with pharmaceutical industries.
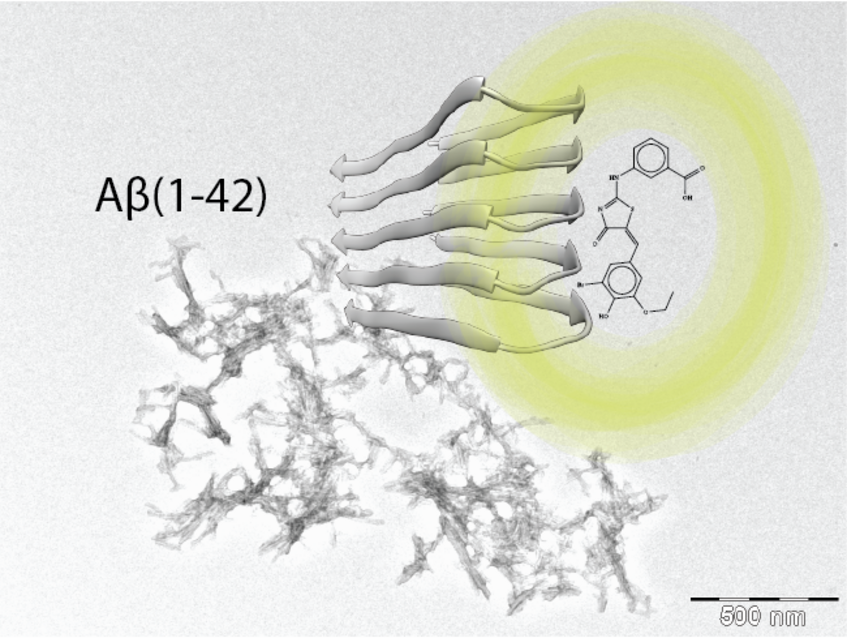
PET Tracer for β-Amyloid Plaques in Alzheimer’s Disease.
PET Tracer for β-Amyloid Plaques in Alzheimer’s Disease.
This project is a collaboration with Prof. Roger Schibli (co-PI), who is the head of the center for radiopharmaceutical science at the Paul Scherrer Institute and the ETH. We aim to develop a PET Tracer for β-Amyloid Plaques in Alzheimer’s Disease. We have already found several molecules that bind to fibrils and inhibit fibrils growth.
NMR2, a method for the fast structure determination of the protein—ligand complex
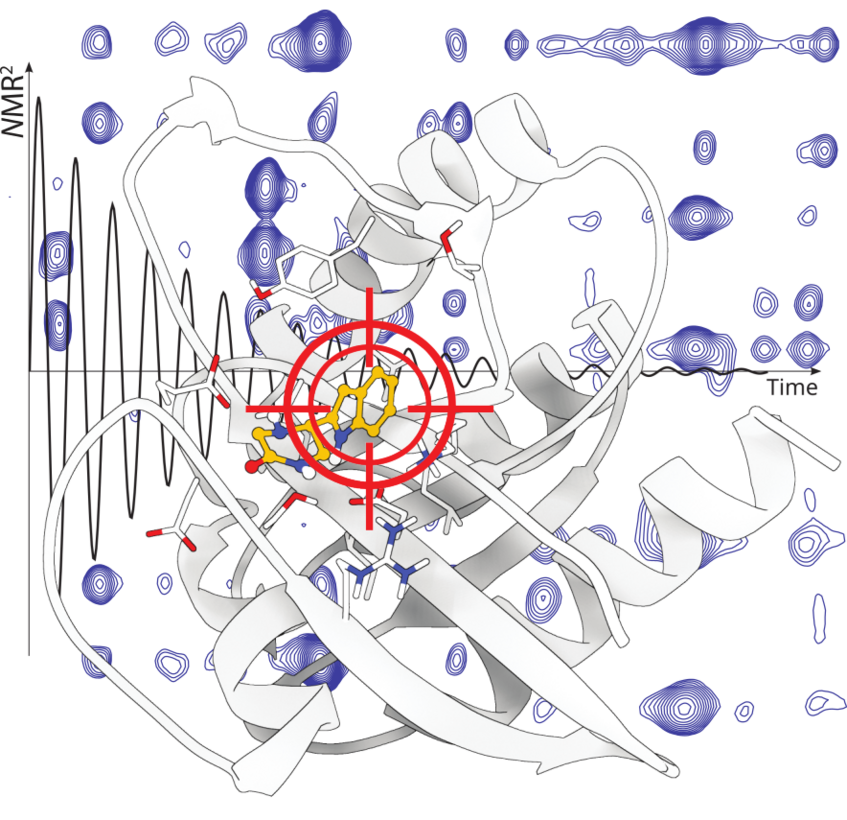
NMR2, a method for the fast structure determination of the protein—ligand complex
Molecular replacement in X-ray crystallography is the prime method for establishing structure-activity relationships of pharmaceutically relevant molecules. Such an approach was not available for NMR. We established a comparable method, called NMR molecular replacement (NMR2) and the scientific community shows high interest for it: “First demonstration that 3D structure determination of a protein–ligand complex can be achieved from solution NMR data fully automatically, including the resonance assignments of the protein.” Nithzche C., Otting G. Current opinion in structural biology, 48, 16-22 (2018).
NMR2 may open a new avenue for the fast and robust determination of the interaction site of protein—ligand complexes at atomic resolution.
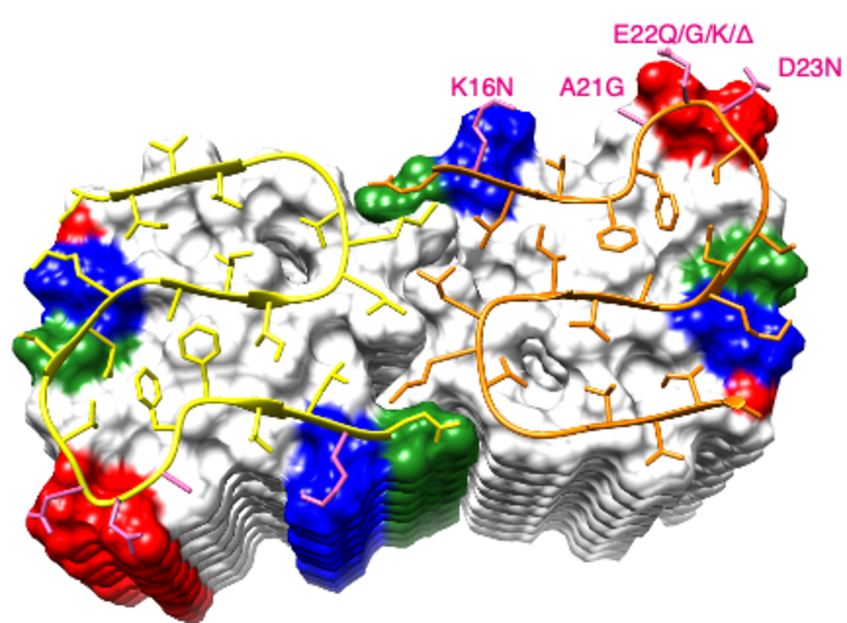
Characterization of disease-relevant Aβ(1-42) amyloid fibrils
Characterization of disease-relevant Aβ(1-42) amyloid fibrils
As one of the pathological hallmarks of Alzheimer’s disease are plaques composed of Amyloid-β (Aβ). Although the two predominant forms, Aβ(1-40) and Aβ(1-42), differ in only two residues, they display different biophysical, biological, and clinical behavior. Aβ(1-42) is the more neurotoxic species, aggregates much faster, and dominates in the plaque of Alzheimer's disease patients. While small Aβ oligomers are believed to be the neurotoxic species, Aβ amyloid fibrils appear to play an important role in disease progression through cell-to-cell transmissibility. We characterized the kinetics of aggregation of Aβ amyloid fibrils by NMR and proposed a model for the aggregation mechanism.
Exact Nuclear Overhauser Enhancement methodology
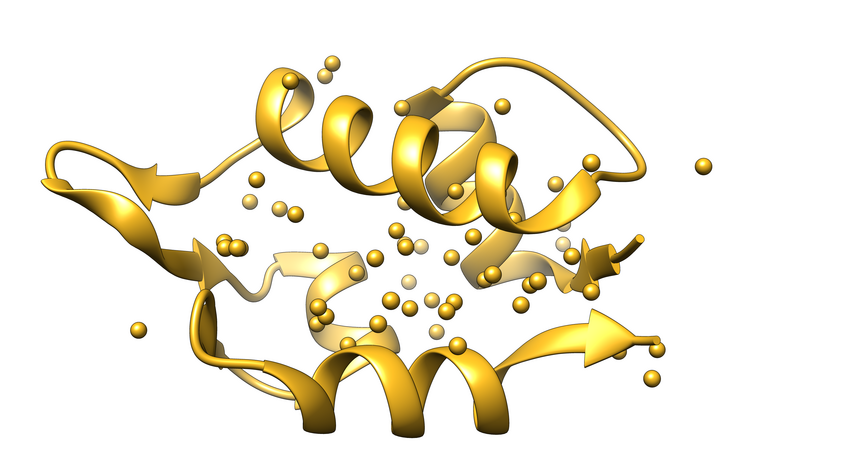
Exact Nuclear Overhauser Enhancement methodology
With the method development on exact Nuclear Overhauser Enhancement (eNOE) precise ensemble-averaged distances between protons with an accuracy of ca. 0.1 Å can be measured. This approach opens an avenue to determine multiple structural states of a protein. We wrote the program called eNORA, that corrects for spin diffusion the NOE data with the full matrix formalism and greatly increases the accuracy of the eNOEs.
INPHARMA
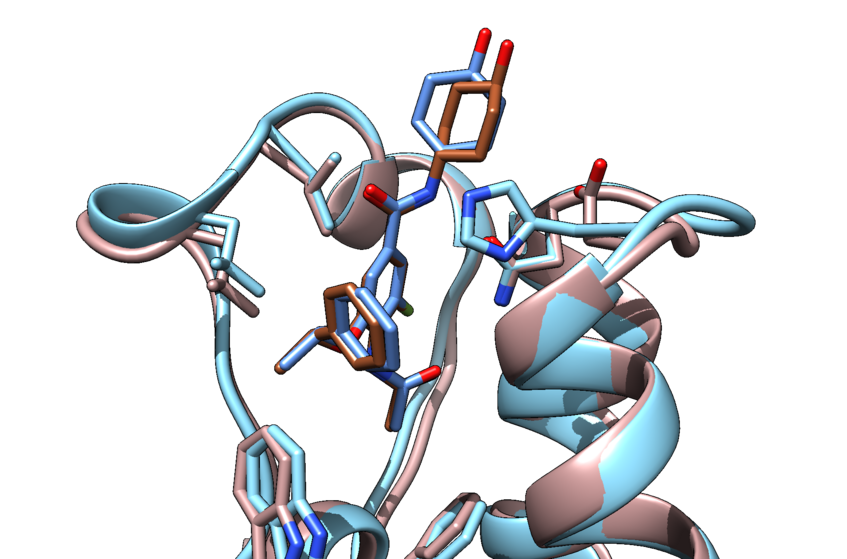
INPHARMA
In the INPHARMA (Inter-ligands Nuclear Overhauser Effect for Pharmacophore Mapping) method, inter-ligand NOEs are used to derive the relative orientation of small molecular binders that compete for the same receptor site. We benchmarked the method using a protein Kinase in complex with fragments. This was a milestone for INPHARMA.
Structural Biology of Autophagy
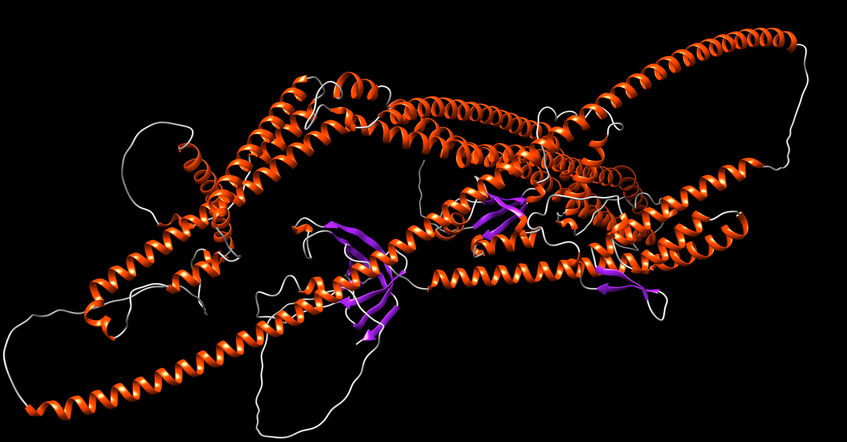
Structural Biology of Autophagy
In collaboration with Sascha Martens' group at the Max Perutz Labs, we are using advanced Nuclear Magnetic Resonance (NMR) methods to analyze the molecular functions of proteins during autophagosome formation.
